
Download PDF
Translations (PDF)
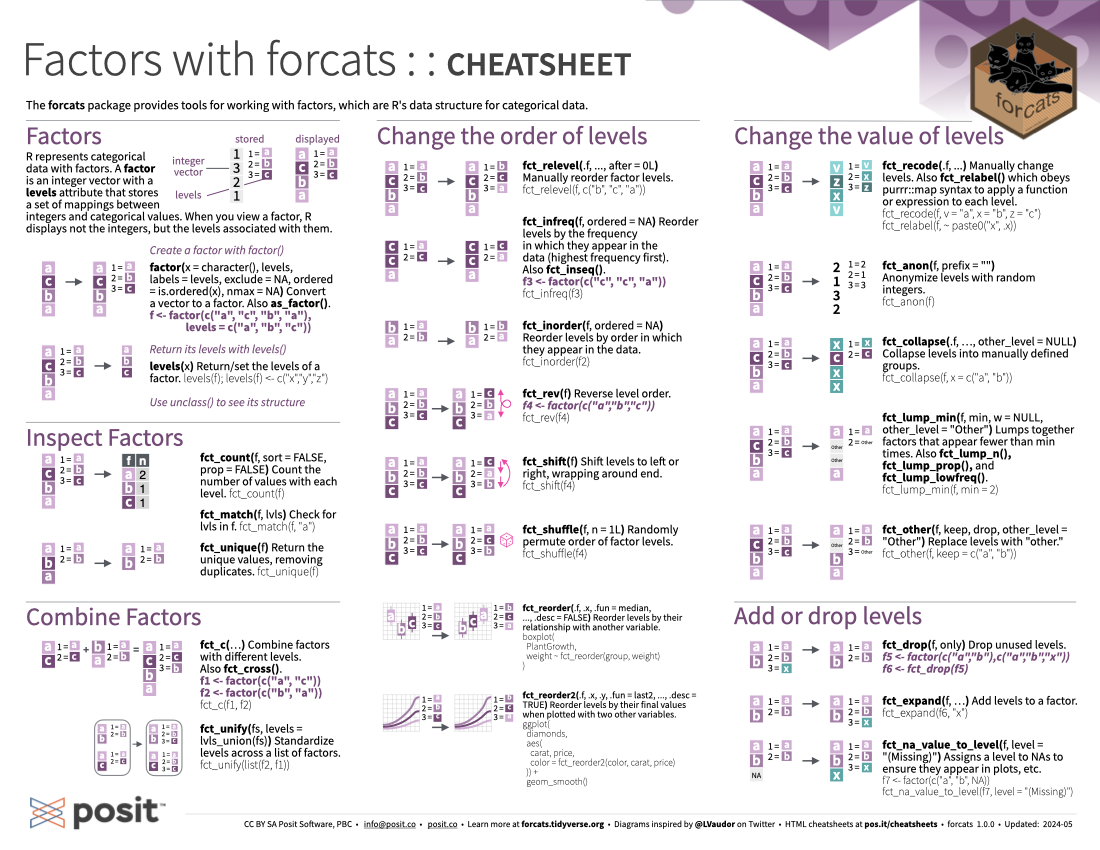
The forcats package provides tools for working with factors, which are R’s data structure for categorical data.
R represents categorical data with factors. A factor is an integer vector with a levels attribute that stores a set of mappings between integers and categorical values. When you view a factor, R displays not the integers but the levels associated with them.
For example, R will display c("a", "c", "b", "a") with levels c("a", "b", "c") but will store c(1, 3, 2, 1) where 1 = a, 2 = b, and 3 = c.
R will display:
[1] a c b a
Levels: a b cR will store:
[1] 1 3 2 1
attr(,"levels")
[1] "a" "b" "c"Create a factor with factor():
factor(x = character(), levels, labels = levels, exclude = NA, ordered = is.ordered(x), nmax = NA): Convert a vector to a factor. Also as_factor().
Return its levels with levels():
levels(x): Return/set the levels of a factor.
Use unclass() to see its structure.
fct_count(f, sort = FALSE, prop = FALSE): Count the number of values with each level.
fct_match(f, lvls): Check for lvls in f.
fct_unique(f): Return the unique values, removing duplicates.
fct_c(...): Combine factors with different levels. Also fct_cross().
fct_unify(fs, levels = lvls_union(fs)): Standardize levels across a list of factors.
fct_relevel(.f, ..., after = 0L): Manually reorder factor levels.
fct_infreq(f, ordered = NA): Reorder levels by the frequency in which they appear in the data (highest frequency first). Also fct_inseq().
fct_inorder(f, ordered = NA): Reorder levels by order in which they appear in the data.
fct_rev(f): Reverse level order.
fct_shift(f): Shift levels to left or right, wrapping around end.
fct_shuffle(f, n = 1L): Randomly permute order of factor levels.
fct_reorder(.f, .x, .fun = median, ..., .desc = FALSE): Reorder levels by their relationship with another variable.
fct_reorder2(.f, .x, .y, .fun = last2, ..., .desc = TRUE): Reorder levels by their final values when plotted with two other variables.
fct_recode(.f, ...): Manually change levels. Also fct_relabel() which obeys purrr::map syntax to apply a function or expression to each level.
fct_anon(f, prefix = ""): Anonymize levels with random integers.
fct_collapse(.f, …, other_level = NULL): Collapse levels into manually defined groups.
fct_lump_min(f, min, w = NULL, other_level = "Other"): Lumps together factors that appear fewer than min times. Also fct_lump_n(), fct_lump_prop(), and fct_lump_lowfreq().
fct_other(f, keep, drop, other_level = "Other"): Replace levels with “other.”
fct_drop(f, only): Drop unused levels.
fct_expand(f, ...): Add levels to a factor.
fct_na_value_to_level(f, level = "(Missing)"): Assigns a level to NAs to ensure they appear in plots, etc.
CC BY SA Posit Software, PBC • info@posit.co • posit.co
Learn more at forcats.tidyverse.org.
Updated: 2024-05.